New Imaging Pipeline Captures Synapse to Whole Brain on One Sample
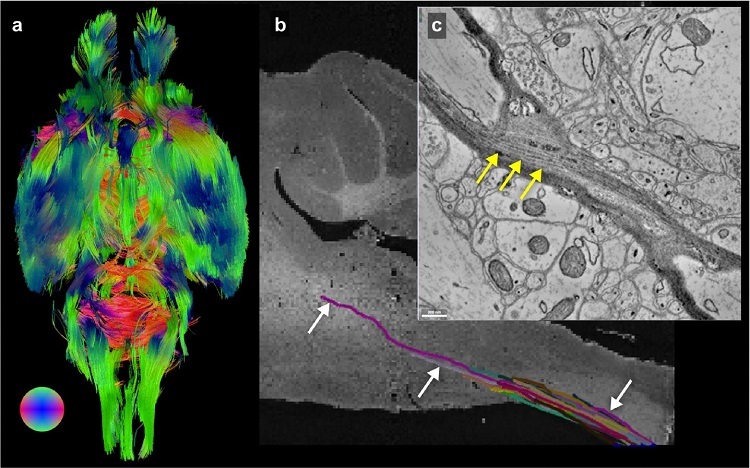
Using three different imaging modalities over five orders of magnitude of resolution, researchers have, for the first time, successfully imaged an entire mouse brain in one go—from the whole brain to individual neurons and tracts.
Previously, researchers struggled to image a single brain sample due to scale-dependent differences in each approach. For example, MRI can image the whole brain to trace neuronal tracts, but the resolution isn’t sufficient to observe individual neurons or their connections. Meanwhile, electron microscopy (EM) can reveal the details of individual synapses, but generates an enormous amount of data, making it challenging to look at pieces of brain tissue larger than a few micrometers. Existing techniques for studying neuroanatomy at the micrometer resolution typically are either two-dimensional or use protocols that are incompatible with MRI or EM imaging.
Like warped pieces of the same puzzle, the incompatibility of the different imaging methods made it impossible to use the same brain tissue for imaging at all scales.
Now, researchers from the University of Chicago and Argonne National Laboratory have pioneered a pipeline using existing methods that can bridge this inconvenient resolution gap to shed a more detailed light on the brain’s multi-scale pathologies.
Argonne is home to the Advanced Photon Source, which is sometimes called the ultimate X-ray microscope. The microscope images in synchrotron-based X-ray tomography (μCT), which can be likened to a micro-computerized tomography (CT) scan. But, this microscope had never previously been used for brain mapping, requiring the team to make a few adjustments to ensure it could simplify the current imaging workflow.
The team employed advances in osmium staining and tissue embedding, worked to extend the field of view of the μCT system with tomographic imaging, and used supercomputers at Argonne to create a parallel algorithm for μCT image reconstruction. Ultimately, the adjustments allowed the researchers to image an entire mouse brain at 1 μm resolution in roughly 6 hours—one of the fastest approaches ever—while outputting only 2 terabytes of data.
Once micron resolution was established, the researchers worked to integrate μCT into a workflow pipeline that would allow multi-modal imaging on a single sample. Using tissue from a single mouse brain, the researchers first used MRI to collect structural images of the entire brain, then turned to μCT to collect CT data before specific regions of interest were identified in the brainstem and cerebellum for targeting with EM.
After months of data processing and image tracing, the researchers were able to use the structural markers identified on the MRI to localize specific neuronal subgroups in designated brain regions, and they could trace the size and shape of individual cell bodies. They could also trace the axons of individual neurons as they traveled through the brain, and could connect the information from the μCT images with what they saw at the synaptic level with the EM.
The researchers say this trifecta of compatible imaging methods could significantly narrow the search for pathological structural change in the brain and lead to improved specificity in MRI through neuroscience. Additionally, it could lead to potential in vivo MRI biomarkers of disease.
“This technique gives us a really clear way to identify changes in the microstructure of the brain when there is a disease or injury present,” said first author Sean Foxley, research assistant professor at UChicago. “Now, we can start looking for biomarkers with the μCT that we can then trace back to what we see on the MRI in the living brain. The X-ray lets us look at things on the cellular level, so then we can ask, what changed at the cellular level that produced a global change in the MRI signal on a macroscopic level?”
In fact, the team is already using the technique to peer into the brains of mice that have been genetically engineered to develop Alzheimer’s disease to see if they can trace the Ab plaques seen with μCT back to measurable changes in MRI scans, especially in early stages of the disease.
While working on this and other projects, the researchers said they are committed to refining the imaging technique.
“The next step is to do an entire primate brain,” said senior author Narayanan “Bobby” Kasthuri, M.D., assistant professor of neurobiology at UChicago and neuroscience researcher at Argonne. “The mouse brain is possible, and useful for pathological models. But what I really want to do is get an entire primate brain imaged down to the level of every neuron and every synaptic connection. And once we do that, I want to do an entire human brain.”
